Test It Out Samples
However, for the deciles we talked about earlier in this thread, I now do not have the specificity nor the sensitivity. I have made an attempt to calculate these figures myself, but since the code does not show how many targets are actually predicted correctly, I am unable to do so.
Do you know how to calculate these measures? First, I notice I made an error in my last block of code in 4. The final command should be -tab TAR prediction, row -. This change will not affect the counts in the cells, but it will change the percentages.
The percentages given by the original code are not the sensitivity and specificity, they are the positive and negative predictive values--which can be important in their own right but are not good indices of prediction accuracy. I'm not sure what you're asking for in 5. Perhaps you mean that you would like to use each decile threshold as a cutoff, that is, get the numbers of predicted and observed targets and non-targets when being in or above that decile is considered a positive prediction.
If that is the case, try this: Code:. Clyde Schechter Excuse me for getting back at my earlier reaction, but I misunderstood the code more specifically, the decile of risk provided in your last reaction.
I thought it basically returned the sensitivity and specificity for different cut-off values. Now I see that it doesn't. In order to avoid typing Code:. You just put this inside a loop: Code:. Clyde Schechter It worked all fine for the ordinary logit model.
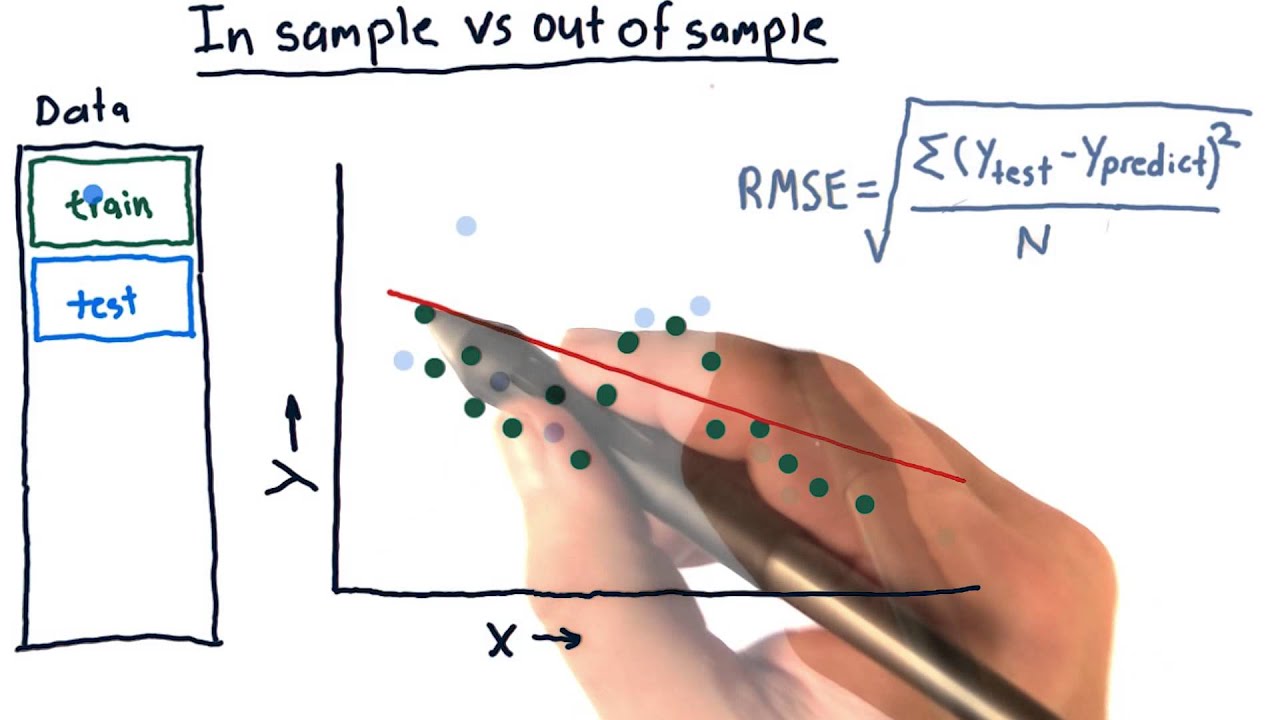
Now a somewhat harder question: is it possible to test the out-of-sample accuracy for the conditional logit model? For this, I think I cannot use the conditional sample, since it only consists of 40 matched pairs, instead of the true sample of ~ firms of which ~40 targets.
This, self-evidently leads to extremely biased results. However, using this method, the other ~ firms would not have predicted probabilities. Do you maybe have any idea on how to assess the out-of-sample prediction accuracy for a conditional logit model?
A conditional logit model doesn't have predicted probabilities, so you can't port them anywhere. That's why your -predict- statement has to use the option -pc it is the probability that each observation in the group will have a positive outcome conditional on there being one and only one positive outcome in the group.
In order to apply this model to the out of sample data, you would need to form matching-groups like those in your -clogit- data, and then do the out-of-sample prediction. But if the "out-of-sample" data consists, as I think you are saying, of the observations for which no match could be found, then this approach is not possible.
Mohamed Elsayed. Dear Clyde Schechter Many thanks for the valuable information you provide in this post and generally in the forum. I have a similar question, please. I was reading a paper in which author was using two models Base Model and Complete Model to predict bankruptcy in one and two years before the bankruptcy using logit model.
To determine which model is better, in one of the tests, the author draws a figure shows the "Prediction Success Rates" I uploaded a pic showing the figure.
The author indicates that the Figure reports the sum of Type I and Type II errors from the base and complete models. The figure shows the sum of Type I errors classifying a bankrupt firm as healthy and Type II errors classifying a healthy firm as bankrupt according to the predicted values computed from each model.
In case if the uploading pic of the figure fails, under the figure, the author provides this definition of what he did to plot that figure: "This figure plots the incidence of Type I errors classifying a bankrupt firm as healthy and Type II errors classifying a healthy firm as bankrupt according to model scores.
Model bankruptcy probabilities are ranked from lowest to highest. For a given percentile, we report the frequency of Type I and Type II errors. For example, at the 50th percentile, we consider the incidence of Type I and Type II errors if all observations with a model bankruptcy probability above the 50th percentile are classified as bankrupt and all others are classified as healthy.
The horizontal axis presents the cutoff point while the vertical axis presents the sum of Type I and Type II errors. Your help will be much appreciated. Thanks in advance, Mohamed. Follow up Clyde Schechter. For your kindly interest. Attached Files. Since you don't provide any data, I will show you an example using mpg as a predictor of foreign in the built-in auto.
Also note that there is no such thing as the 0th percentile, nor the th. Hi Clyde, Many thanks for advising me on this. In terms of data, I should duplicate the figure I showed in my previous post using any data.
But, let me explain the sample, the variable and model I should use. The sample contains a number of firms that go bankrupt and healthy firms during this period, so it's panel or pooled dataset. Base Model is nested in the Complete Model.
I already run the multi-period logit and get the coefficients by using the code " logit y x1 x2 x3 i. year i. indusrty" for the base model and "logit y x1 x2 x3 x4 i. indusrty" for the complete model.
Now it is required from me to plot a similar plot to the one I indicated in my previous post. Please note that to predict bankruptcy for year and two years in advance like the paper in my post, I used lag1 and lag2 for independent variables.
for example logit y l. Below, we are doing this process in R. We are first simulating two samples from two different distributions. These would be equivalent to gene expression measurements obtained under different conditions.
The resulting null distribution and the original value is shown in Figure 3. FIGURE 3. The original difference is marked via the blue line. The red line marks the value that corresponds to P-value of 0. After doing random permutations and getting a null distribution, it is possible to get a confidence interval for the distribution of difference in means.
We can also calculate the difference between means using a t-test. Sometimes we will have too few data points in a sample to do a meaningful randomization test, also randomization takes more time than doing a t-test.
This is a test that depends on the t distribution. The line of thought follows from the CLT and we can show differences in means are t distributed.
There are a couple of variants of the t-test for this purpose. If we assume the population variances are equal we can use the following version.
We can calculate the quantity and then use software to look for the percentile of that value in that t distribution, which is our P-value. For this test we calculate the following quantity:. Luckily, R does all those calculations for us.
Below we will show the use of t. test function in R. We will use it on the samples we simulated above. A final word on t-tests: they generally assume a population where samples coming from them have a normal distribution, however it is been shown t-test can tolerate deviations from normality, especially, when two distributions are moderately skewed in the same direction.
This is due to the central limit theorem, which says that the means of samples will be distributed normally no matter the population distribution if sample sizes are large. We should think of hypothesis testing as a non-error-free method of making decisions.
Similarly, we can fail to reject a hypothesis when we actually should. And we usually want to decrease the FP and get higher specificity. And, again, we usually want to decrease the FN and get higher sensitivity.
More powerful tests will be highly sensitive and will have fewer type II errors. For the t-test, the power is positively associated with sample size and the effect size. The larger the sample size, the smaller the standard error, and looking for the larger effect sizes will similarly increase the power.
We expect to make more type I errors as the number of tests increase, which means we will reject the null hypothesis by mistake. However, if we make tests where all null hypotheses are true for each of them, the average number of incorrect rejections is There are multiple statistical techniques to prevent this from happening.
These techniques generally push the P-values obtained from multiple tests to higher values; if the individual P-value is low enough it survives this process. However, this is too harsh if you have thousands of tests.
Other methods are developed to remedy this. This gives us an estimate of the proportion of false discoveries for a given test. To elaborate, p-value of 0. An FDR-adjusted p-value of 0. The FDR-adjusted P-values will result in a lower number of false positives.
Within the genomics community q-value and FDR adjusted P-value are synonymous although they can be calculated differently. In R, the base function p. adjust implements most of the p-value correction methods described above.
For the q-value, we can use the qvalue package from Bioconductor. Below we demonstrate how to use them on a set of simulated p-values. The plot in Figure 3. FDR BH and q-value approach are better but, the q-value approach is more permissive than FDR BH.
In genomics, we usually do not do one test but many, as described above. That means we may be able to use the information from the parameters obtained from all comparisons to influence the individual parameters. For example, if you have many variances calculated for thousands of genes across samples, you can force individual variance estimates to shrink toward the mean or the median of the distribution of variances.
This usually creates better performance in individual variance estimates and therefore better performance in significance testing, which depends on variance estimates.
How much the values are shrunk toward a common value depends on the exact method used. These tests in general are called moderated t-tests or shrinkage t-tests. Bayesian inference can make use of prior knowledge to make inference about properties of the data. In a Bayesian viewpoint, the prior knowledge, in this case variability of other genes, can be used to calculate the variability of an individual gene.
Below we are simulating a gene expression matrix with genes, and 3 test and 3 control groups. Each row is a gene, and in normal circumstances we would like to find differentially expressed genes. In this case, we are simulating them from the same distribution, so in reality we do not expect any differences.
Listen to Check it out. Royalty-Free sound that is tagged as acappelas, funk/soul, hip hop, and vocals. Download for FREE + discover 's of sounds Out-of-sample testing is used to evaluate the performance of a strategy on a separate set of data that was not used during the development and See all of “Check It Out” by Nicki Minaj & canadian24houropharmacy.shop's samples